By Chris Gignoux and Mike Macpherson
It should be no surprise that in general, we are more genetically similar to our neighbors than to people living far away. The reason is fairly simple – until recently in human history it was fairly rare for people from widely separated geographic regions to even meet, much less reproduce.
This pattern, known as isolation-by-distance, has been observed in a number of studies over the past several decades. This week, it has been confirmed in Europe by the largest study of its kind to date.
The researchers produced a two-dimensional map, like the one below, that preserves the genetic similarities between individuals as far as possible; in other words, the closer two dots (people) are on the map, the more closely related they are genetically.
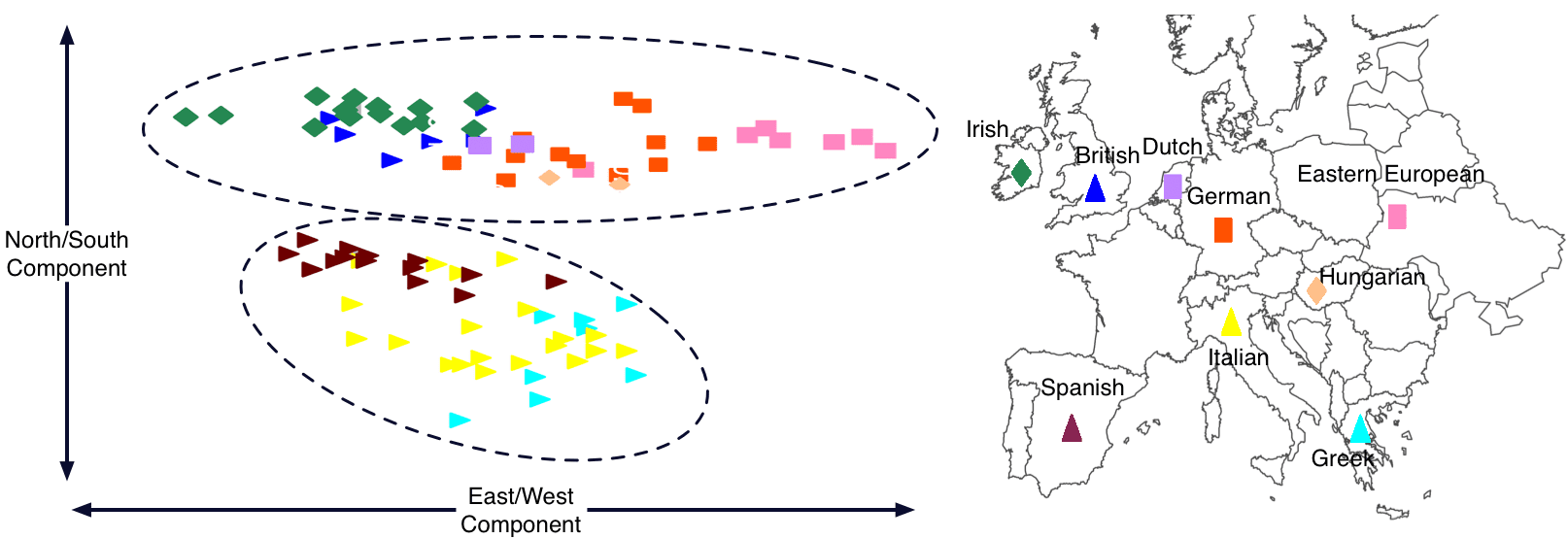 A two-dimensional genetic similarity map of Europeans shows the northern and southern clusters. Each colored symbol in the plot on the left represents a person’s genotype. Note the similar placement of symbols on the plot to the left and the geographic legend to the right. Adapted from Tian et al., Plos Genetics, (2008).
A two-dimensional genetic similarity map of Europeans shows the northern and southern clusters. Each colored symbol in the plot on the left represents a person’s genotype. Note the similar placement of symbols on the plot to the left and the geographic legend to the right. Adapted from Tian et al., Plos Genetics, (2008).
In the figure above, each individual was labeled with their country of origin after the mapmaking procedure was run. If Europe were genetically homogeneous, you would expect the different nationalities to appear in a jumble. Instead, they separate into clusters that remarkably recapitulate the geography of Europe.
Northern vs. Southern Europe
Even though Europe has been occupied for only a relatively short time compared to other parts of the world, different populations within the continent have had time to differentiate from one another. Scientists have known for a long time that certain traits, like lactase persistence and light-colored eyes and hair are more common in northern than in southern Europe. Likewise, there are certain diseases such as sickle cell anemia that, although rare across Europe, are found more in the south than in the north. Height and skin color also vary from northern to southern Europe: both vary gradually with latitude rather than in quick jumps.
Early genetic studies (such as those in the landmark population genetics text History and Geography of Human Genes) showed that this north-south cline was also a genetic one: even though Europeans of different nationalities did not fit into simple clusters, there was an overarching north-south difference. Newer studies have increased the number of people typed, and the number of markers, to approach the genome-wide level of hundreds of thousands of SNPs we use here at 23andMe – which brings us to this week’s paper.
A summary of genome-wide findings
The Lao et al. study out this week obtained genotypes from more than 2,500 individuals of known European ancestry. Each of the genotypes consists of about half a million SNPs typed on the Affymetrix 500K, a chip similar in size to the Illumina 550K used here at 23andMe. They confirm the findings of several recent but smaller European studies (Seldin et al, PLoS Genetics (2006); Bauchet et al, AJHG (2007); Tian et al, PLoS Genetics (2008); Price et al, PLoS Genetics (2008); Paschou et al, PLoS Genetics (2008)), namely:
- Overall, all SNPs and Europeans are very genetically similar.
- A small set of SNPs does allow European populations to be distinguished—at least when used among people whose ancestors are all from the same part of Europe—and they are surprisingly effective.
- Most of the genetic variation in Europe is found along the north-south axis, which is consistent with archaeological knowledge. The next most prominent axis of genetic variation runs roughly east-west.
- More isolated populations tend to exist at the extremes of these plots. In this current paper, the Finns are the only nationality utterly distinct from the rest of the European samples. The Finns speak a different language from much of the rest of Europe.
There’s plenty of action in the blogosphere on this one. For more discussion, check out dienekes’ anthropology blog, anthropology.net, gene expression, and genetic future.