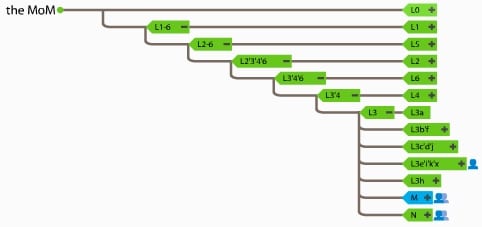
It’s springtime for 23andMe’s Maternal Line tree!
Each one of us can trace our ancestry back through our mother, our mother’s mother, her mother, and so on. And there is one piece of our DNA that has been inherited along that exact, maternal line.
By examining mutations in this segment of DNA, called mitochondrial DNA because it resides in parts of the cell called mitochondria, we can learn about our line of maternal ancestors – in particular, where these ancestors came from. Sets of similar, closely related maternal lines are called “haplogroups” and are given labels such as “J1c3” or “X2b”. Numerous researchers around the world study these maternal haplogroups to better understand where they originated and where people with the haplogroups have migrated over the course of thousands of years.
23andMe interprets the haplogroups for customers based on the publications of these researchers.
All maternal haplogroups are themselves related; the relationships are represented as a “maternal line tree”. 23andMe’s version of that tree has just experienced both pruning and new growth via an extensive update. With these changes, a customer’s particular branch (maternal haplogroup) may or may not have changed, but, overall, this new tree is far bushier, with over 1500 different haplogroup assignments possible, and over 870 haplogroups actually assigned to customers.
Why update the haplogroups?
Maternal haplogroups are of greatest interest if (a) they have a known history or (b) they enable people to discover others who are related to them. Scientists are continually revising the tree that indicates which mutations define each maternal haplogroup, and how all the haplogroups are related. The more up-to-date the tree used for haplogroup assignments is, the more accurate and more meaningful those assignments are. Up-to-date haplogroups enable our customers to answer questions such as the following:
a) Which haplogroup histories are relevant to me?
Your maternal haplogroup and the maternal haplogroups of your ancestors provide insights into your ancestry. Let’s say your haplogroup under 23andMe’s previous tree was H2. The known history for haplogroup H2 is very general; women with this haplogroup survived and reproduced very successfully in Europe during the Last Ice Age. Let’s say your haplogroup under 23andMe’s new tree is now H2a2. While H2 is common throughout all of Europe, research has shown that H2a is concentrated near the Caspian Sea. The history for H2a2 has yet to be written, but eventually we’ll understand, possibly through 23andMe’s research, the origin and migration history of this haplogroup that branches off from H2a.
b) To whom am I most similar?
Who are your maternal line cousins? In general the higher resolution the haplogroup, the more meaningful the sharing of a haplogroup. For example, if your haplogroup is H2, you share that haplogroup with over 600 23andMe customers; on the other hand, if your assignment is H2a2b, you share that with just over 60 customers. The common ancestor of everyone with H2a2b lived far more recently than the common ancestor of everyone with H2. People with a more specific haplogroup can begin to ask, where are we all from? Where did our common ancestor live, and when?
23andMe also aims for consistency with other genetic services. If all such services are using the same haplogroup definitions, then maternal haplogroups are comparable across services.
Steps in the update
In the summer of 2007 when 23andMe developed its initial Maternal Line feature there was no publicly available, up-to-date tree. Therefore 23andMe used information from various sources to create a tree as up-to-date as possible and to assign haplogroups.
Now there is a publicly available tree maintained by scientist Mannis van Oven of the Netherlands, with help from other geneticists and genetic genealogists (including 23andMe customers!). They update the tree regularly, and we used this tree (Build 7) as a guide for updating our Maternal Line feature. We recalculated every customer’s haplogroup and otherwise modified the feature to be consistent with the tree of Phylotree.org.
Many customers will see the same maternal haplogroup as in the past. Thousands, however, will see something new.
Customer haplogroups have changed for two main reasons: (1) because of new definitions of the haplogroups, and (2) because of improved SNP results.
Some customers will see a higher resolution haplogroup.
For example, over 100 customers see a change from H4a to H4a1a. Other customers will see a major change in their haplogroup name, even though the history of that haplogroup has not changed. For example, some customers who had an H5a assignment will now see H36. Others who had PreHV1 will now see R0. Some customers will see a less dramatic change: all customers with a J1a haplogroup will now see a J1c haplogroup. Many of these changes simply reflect relabeling of the tree by van Oven and other scientists.
The most common change is the removal of a “*” from a haplogroup label. The “*” is used to indicate that a person’s haplogroup cannot be assigned to a more specific subhaplogroup (for example H* means that a person is not in H1, H2, etc). As the maternal haplogroup tree grows, the “*” will necessarily change in meaning, so 23andMe has decided to do away with it and instead use a system in which we just assign people to the most specific haplogroup possible.
So if you are definitely in haplogroup H, but not a part of H1, H2, etc , then you will simply be assigned to haplogroup H.
In some cases a haplogroup has been split. So some people with T2b2 will now have T2b4, while others continue to have T2b2. Sometimes customers will have less detailed haplogroups. For example, a few hundred customers go from T2b2 to T2b because T2b2 was redefined.
Send us your feedback!
With over 870 distinct haplogroups assigned to more than 35,000 customers, we anticipate making further refinements. If you have information regarding your mtDNA that is inconsistent with our update, please let us know at [email protected]!
Reference: van Oven M, Kayser M. 2009. Updated comprehensive phylogenetic tree of global human mitochondrial DNA variation. Hum Mutat 30(2):E386-E394. http://www.phylotree.org. doi:10.1002/humu.20921



