With the holiday season upon us, 23andMe is sprucing up its paternal haplogroup assignments!
With 23andMe population geneticist and Y-chromosome expert David Poznik at the lead, we’ve updated our Haplogroups 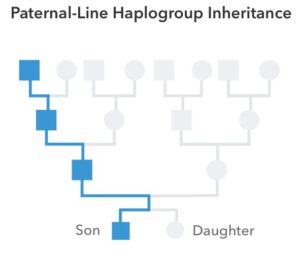 Report and paternal haplogroup assignemtns. The update reflects significant developments in the field over the past few years. We’re also excited to introduce yHaplo™, our new open-source software for researchers.
Report and paternal haplogroup assignemtns. The update reflects significant developments in the field over the past few years. We’re also excited to introduce yHaplo™, our new open-source software for researchers.
Major Refinements to the Y-Chromosome Tree
Each generation, fathers pass down copies of their Y chromosomes to their sons. Small variations arise over time and accumulate in patterns that uniquely mark individual paternal lineages. To trace the evolutionary history of these lineages, scientists study DNA sequence differences between and among modern populations. Using this information they have built a “tree” that shows how global Y chromosomes relate to one another.
However, our understanding of the Y-chromosome tree had, for many years, been limited by our incomplete knowledge of Y-chromosome diversity. Because paternal haplogroup names reflected the structure of the tree, each new insight required renaming haplogroups. This in turn made it difficult to interpret paternal haplogroup assignments from one year to the next.
Recent research, including a study published in Nature Genetics, has drastically refined the structure of the tree. For that work, David and an international team of 42 scientists used complete Y-chromosome sequences from around the world to carry out the largest-ever study of genetic variation within the human Y chromosome (Poznik et al.). This research identified more than 65,000 Y-chromosome genetic variants. The study vastly increases our understanding of the tree and sets a new standard for tracing male lineages through migrations that have occurred over the millennia of human history.
What’s Changing
Male customers on the new 23andMe website experience can expect a couple of changes to their paternal haplogroup assignment with this update. Female customers may also see changes to the paternal haplogroup assignments of male relatives and friends in other parts of the website.
First, we have substantially updated our Y-chromosome tree to reflect the work of the International Society of Genetic Genealogy (as of January 4, 2016).
In most cases, the updated haplogroup assignments are equivalent to previous assignments or differ only slightly. However, since much more is now known about the tree, we can provide more information about an individual haplogroup’s history and how it relates to others.
The second major update is a change to the naming system we use to report paternal haplogroups. Until recently, the convention was to use an often lengthy series of letters and numbers indicating the path of branches from the most recent common ancestor of all men to each haplogroup.
The problem is that these names changed from year-to-year as the tree was refined, making it difficult to know from the name alone which haplogroup male customers actually carry.
To reduce confusion, we have moved to a system of shorter and more stable names. Each name uses a letter to identify the major branch of the tree and the name of a genetic marker unique to a specific haplogroup. For example, if we previously reported your paternal haplogroup as “Q1a3a,” we now report it as “Q-M3,” indicating that your Y-chromosome lineage belongs to a subgroup of haplogroup Q that bears the M3 marker. Because this new representation focuses on a specific informative marker associated with your haplogroup, it will be much more stable over time.
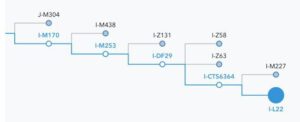
A small section of the updated Y-chromosome tree illustrating the marker-based haplogroup naming convention. The structure of the tree was aggregated from the literature by the International Society of Genetic Genealogy.
For more information on the changes coming to the haplogroups report, visit 23andMe’s customer care page, here.
yHaplo™, a New Open-Source Research Tool
The paternal haplogroup update doesn’t end with the tree. As a member of the research team at 23andMe, Poznik has developed a new algorithm to rapidly and robustly identify Y-chromosome haplogroups in very large samples. In addition, he has implemented the algorithm as the yHaplo software package.
This software is very flexible; it runs on full Y-chromosome sequences and on smaller sets of genotyped markers. Furthermore, it is easy to incorporate updates as researchers around the world continue to gather data and learn more about the Y-chromosome tree.
At 23andMe, we’re using this software to provide paternal haplogroup assignments to our customers.
As we believe the yHaplo software package can be an extremely useful tool to help drive research, we have made it available under a custom open-source software license for non-commercial research use. To learn more about yHaplo, read our white paper or head to the code repository!



