This guest post is by Roy King, who is a professor of psychiatry at Stanford University and a research colleague of Stanford geneticist and 23andMe scientific adviser Peter Underhill. Roy and Peter have been using genetics to trace the spread of agriculture from the Near East to Europe.

The question of how agriculture first arose and spread in Europe has perplexed archaeologists and geneticists alike for decades. Evidence for farming communities in Greece, Crete, the Balkans, Southern Italy, and Mediterranean France and Spain first appears in the archaeological record eight to nine thousand years ago with the appearance of domesticated wheat, barley, sheep, goats and cows, and a related culture featuring pottery and anthropomorphic figurines. Before this period, most of the indigenous people of Europe fished, hunted small game or foraged for their livelihood.
But how did farming get to Europe in the first place? We know that 2,000 years before agriculture hit Europe, just after the end of the Ice Age, people living in the Near East had already developed farming, with the domestication of wild species of grasses, goats and cattle likely beginning in the fertile river valleys in present-day northern Iraq, Syria and southeastern Turkey. Near Eastern farmers also settled in villages and produced pottery and ceramic human figurines similar to the ones later found across Mediterranean Europe.
So did the first farmers of the Near East hop into boats with their domesticated plants, animals and artistic motifs and colonize Mediterranean Europe? Or did native Europeans learn about farming through trade with the Near East and decide to adopt this agricultural economy too?
Genetic studies are starting to provide answers to this enduring question. Not only do genetic studies of sheep, goats, cows and wheat demonstrate that the European varieties are subtypes of the species found in the Near East, but data from the human Y chromosome suggest that people currently living in Southern Europe are descended in large part (at least through the paternal line) from the Near East. This suggests, at least for the Mediterranean areas of Europe, that the most probable scenario is that Near Eastern farmers did actually move into Europe, bringing farming with them.
23andMe has a Y chromosome marker on its custom chip, rs34126399, which captures the spread of agriculture from the Near East to Europe. The G state at rs34126399 is found in most individuals carrying paternal haplogroup J2a, whose origin can ultimately be traced to Turkey 15,000 to 20,000 years ago. Southern Turkey is the likeliest source for the initial domestication of wheat.
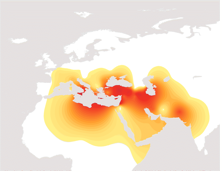
In the millennia after its appearance in Turkey, rs34126399G then experienced a population growth and spread westward into the eastern Mediterranean and eastward into Mesopotamia, Iran and northern Egypt. The map to the right shows the present-day distribution of rs34126399G in Europe and the Middle East.
In addition to its high frequency in the Near East, rs34126399G is present in 10 to 25% of the population of southern Greece and Italy, Crete, Sicily, Bulgaria and Romania. The concentration is an indication of the early spread of farming to these regions, where the climate of wet winters and hot dry summers suited the varieties of winter wheat that were first domesticated.
Since the initial movement of people associated with the origins of agriculture nearly 10,000 years ago, there has been persistent contact and trade throughout the Mediterranean. Just think of the Mediterranean diet – rich in olive oil, goat cheese, and red wine – all of whose origins can be traced to these regions. One archaeologist described the Mediterranean Sea figuratively as a giant bathtub that for millennia has permitted easy seafaring transit from one area to another. The genetic evidence appears to support that analogy.