Starting in January with the launch of the ambitious Precision Medicine Initiative in the United States, 2015 finished with a  heated debate around the appropriate use of technology that allows scientists to edit the human genome.
heated debate around the appropriate use of technology that allows scientists to edit the human genome.
Those two big stories bookended the year showing both the promise and peril of genetics.
2015 triggered a furious pace in new genetic insights with all the potential they hold. But with many of these advances comes the weight of new issues to tackle as well. As we head into 2016, we’re taking a moment to look back at some of the highlights of the year including 23andMe’s contributions to the science and discussion around genetics.
Transforming Medical Research
The Precision Medicine Initiative announced by President Obama in January could help change the face of medicine forever. The effort to sequence a million Americans will depend on the willingness of people to participate in research meant to, as Obama said, “bring us closer to curing diseases like cancer and diabetes, and to give all of us access to the personalized information we need to keep ourselves and our families healthier.” It’s the kind of people-powered research model that 23andMe pioneered; one that is about more than just collecting data, but depends on empowering consumers and engaging them in their own health.
A Banner Year for 23andMe
While it wasn’t a scientific breakthrough, syndrome carrier status report was a huge step for direct-to-consumer health testing in general. It was the first time the US regulatory agency gave such authorization, and it’s laid the groundwork for how to return actionable health results to consumers in a regulated environment. It also was a highpoint in a long list of highpoints for 23andMe, that included launching a Therapeutics group and introducing a newly imagined product experience for customers that included health, ancestry and traits reports and the first-ever genetic test with reports that meet FDA standards.
syndrome carrier status report was a huge step for direct-to-consumer health testing in general. It was the first time the US regulatory agency gave such authorization, and it’s laid the groundwork for how to return actionable health results to consumers in a regulated environment. It also was a highpoint in a long list of highpoints for 23andMe, that included launching a Therapeutics group and introducing a newly imagined product experience for customers that included health, ancestry and traits reports and the first-ever genetic test with reports that meet FDA standards.
The People of the British Isles
History can be found in our DNA, and new research by scientists at Oxford University found distinctive regional differences in the tight confines of the British Isles. The study, led by Sir Walter Bodmer, also found that there is no single “Celtic” genetic group. Instead the study found that people in the Celtic regions of the UK – Scotland, Northern Ireland, Wales and Cornwall – are among the most genetically diverse in all the United Kingdom. In addition, the researchers estimate that the genetic contributions from Anglo-Saxon migrations – Germanic people who came into the coastal lowlands of the British Isles in the 5th and 6th centuries– to be just under half among people in the south east of England. As with studies done by 23andMe looking at African ancestry in the United States, this fine scale genetic map of the British Isles illuminates historical patterns of migration and distinct regional differences. It even reveals clusters of populations that show similar geographic locations of tribal groupings and kingdoms dating back to the 6th century.
That Dress

IMAGE: TUMBLR / SWIKED
Scientists at 23andMe ask a lot of questions. It’s their job, and it informs the kind of innovative research 23andMe pioneered. The information we get from people who participate in research is combined with genetic information that feeds into 23andMe’s powerful and novel research model. It’s allowed our scientists to quickly find genetic insights related to human traits, inheritance and disease. A great recent illustration of that came earlier this spring when 23andMe computational biologist Fah Sathirapongsasuti, PhD, tapped into the flurry of conversation created by The Striped Dress, where people saw distinctly different colors: blue and black, or white and gold. In a matter of a few days 23andMe was able to survey about 25,000 customers interested in participating in the study, and make some interesting findings. While there was no clear genetic association, Fah found a small effect size in a gene associated with light perception – ANO6. But it was non-genetic factors – namely a person’s age – that seemed to play the biggest role in why some people saw one pair of colors and others saw something else.
Got Lactase?
Nutrient-rich with plenty of vitamin D for the sun-starved in Northern Europe, milk was a kind of superfood of the Bronze Age. But new research published in the spring indicates that it took awhile before Europeans could digest milk, and that trait appears to have been introduced by the Yamnaya, a group of people who migrated into Northern Europe from an area near the Altai Mountains in what is now central Russia.
You can read more here.
A Little Buggy
Some people are just tastier to mosquitoes than others, and that has to do with genetics, according to a twin study done by researchers at the London School of Hygiene & Tropical Medicine. Understanding these differences could help in fighting the spread of devastating mosquito born illnesses like malaria and dengue fever.
Optimism about Research into Depression
For the first time ever researchers identified two genetic variants associated with major depressive disorder. In a study published in Nature over the summer, scientists at University of Oxford found the genetic variants and were able to replicate their findings. Although previous studies have found genetic associations, unlike with these findings those failed to stand up to further scrutiny. One of the variants identified in this study is near the gene SIRT1, which plays a role in mitochondria, the energy producing organelles of the cell. The findings may help identify drug targets to better treat depression, which affects more than 350 million people.
Innovation in Diagnosing Parkinson’s
A team of researchers has developed a simple, non-invasive approach to diagnosing people with Parkinson’s disease, an illness that is typically very difficult to detect. The model uses a combination of such things as genetic data, family health information, age, and biomarkers like the ability to smell to make a diagnosis. The researchers believe that with some refinements the tool could help in screening and diagnosing individuals for the condition and help in differentiating Parkinson’s disease from other neurological conditions.
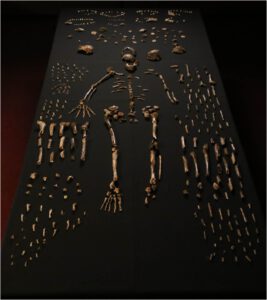
Composite skeleton of Homo nadeli along with other bones found at the Rising Star Cave in South Africa. Photo: Lee Berger, Wits, photographed at Evolutionary Studies Institute
More Cousins
Fossils found in a South African cave could be that of a brand new human ancestor, dubbed Homo naledi. The species dates back two million years, and pushes back the age of the genus Homo – which includes modern humans – at least 400,000 years earlier than previously thought.
Gene Editing
Few breakthroughs have stimulated more discussion in genetics than the innovative technique to “edit” specific bits of DNA. Called Crispr-Cas9, it offers a simple and quick way of snipping out a sections of DNA and stitching the DNA back together again. In so doing it could be used for all sorts conventional and gene therapies. Yet while the gene editing tool offers intriguing possibilities for “correcting” genetic defects that trigger disease or inherited conditions, it raised huge moral and ethical issues. While the technology has raced ahead, sorting out all the issues it raises has lagged behind. In December scientists from around the world met to discuss the implications in Washington DC and agreed to put a halt to editing of the human genome until clearer lines and standards could be agreed upon although research continues.



